A) PDB Input Selection
Please enter either a single PDB code of interest (see the Help page for more details).
Error: PDB with < 10 or > 600 c-alpha atoms not allowed
B) NMA parameters
Use the drop down menu to change force field. The C-alpha force field is recommended for most applications.
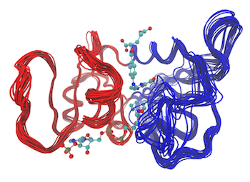
Input Structure Visualization
Input PDB Read Log
Normal mode analysis with Bio3D
This Bio3D WebApp provides a rapid and rigorous tool for normal mode analysis of protein structures. Options include multiple popular elastic network models (ENMs), as well as enhanced analyses including residue fluctuations, mode visualization, dynamic cross-correlations, and overlap analysis.
Start by entering a PDB code of interest then proceed by navigating through following the NEXT buttons.
Results
Plot options
Plot options
B-factors
B-factors
Overlap analysis
Plot options
Overlap values
Cumulative overlap values
References
Force fields
calpha: Hinsen, K. et al. (2000) Chemical Physics 261 25-37
anm: Atilgan, A.R. et al. (2001) Biophysical Journal 80 505-515.
sdenm: Dehouck Y. & Mikhailov A.S. (2013) PLoS Comput Biol 9 :e1003209.
reach: Moritsugu K. & Smith J.C. (2008) Biophysical Journal 95 1639-1648.
pfanm: Yang, L. et al. (2009) PNAS 104 .